In the previous article in this series, we gave a basic introduction to scientific computing with Chapel by porting over the 4th step of the CFD Python: 12 steps to Navier-Stokes tutorial. The focus on a simple 1D example allowed us to introduce Chapel’s syntax as well as concepts like arrays, ranges, and forall loops.
In this post, we’ll use Steps 9 and 10 of the tutorial (which cover Laplace’s and Poisson’s Equations, respectively) to introduce some more advanced topics. We’ll then discuss a performance comparison with Python and C++/OpenMP to show that Chapel is able to deliver readable Python-like code while matching the performance of well-known HPC tools like OpenMP. Note that the code discussed in this article primarily follows the structure of Step 10 while also drawing some inspiration from Step 9.
In the next couple of posts, we’ll build off of the concepts introduced here to create a distributed, multi-node version of Step 11 — a full Navier-Stokes cavity-flow simulation. And to continue our performance investigation, we’ll also compare the Chapel code’s performance with an equivalent code written with C++/MPI/OpenMP.
Poisson’s Equation in Chapel
In the context of the Navier-Stokes Equations, Poisson’s equation is used to describe how a fluid’s pressure is distributed as a function of its momentum. It is also used in other scientific domains to describe things like electric potential or gravitational potential in terms of charge or mass distributions. As such, a performant and easy-to-read Poisson solver like the one we’ll discuss here can be useful in a variety of applications.
Mathematical Background
For our purposes, Poisson’s Equation can be written as follows, where is a scalar quantity representing a fluid’s pressure as a function of space, and is a scalar source term, which in the incompressible Navier-Stokes context can be derived from the fluid’s momentum:
For the 2D case, it can be written as:
Notice that there is no time component. This means that the equation is not describing how the fluid’s pressure evolves over time. Rather, it is simply describing ’s shape by saying that at any point in space, the sum of its partial 2nd derivatives (curvature) in each direction should be equal to the source value at that point in space.
As a result, the finite difference method will be applied in a slightly different manner than in the previous post. Instead of using a stencil operation to compute the next time step, we’ll repeatedly apply a stencil to until we have an approximate solution that satisfies this discretization of Poisson’s Equation:
Note that the and subscripts represent discrete and coordinates.
Because we’re not iterating for a predetermined number of discrete time steps, we’ll need a way of deciding how many times to apply the stencil operation. We can either iterate until the difference in from step to step is very small, meaning that the algorithm has asymptotically approached the solution (covered in Step 9), or we can iterate for some large number of steps, assuming that a sufficiently accurate solution will have been computed by that point (covered in Step 10). In this article, we’ll implement a code that can use either termination condition.
Source and Boundary Conditions
From Step 10 of the tutorial, we’ll use the following value for that should create a peak and valley in our solution around the two specified points:
And to keep things interesting, we’ll take the boundary conditions from Step 9, which includes a linear Dirichlet boundary condition on the right wall that we’ll be able to see in our solution:
Simulation Setup
To start things off, we declare some constants at the beginning of our program that we’ll use to implement the remainder of the simulation:
|
|
These mostly follow the values from the tutorial, but we’ve also included a couple extra things:
- There is a configurable constant,
makePlots, to control whether the solution gets plotted with an external script. SourceMagwill control the magnitude of the peak and valley in the source term.
Additionally, a configurable param is declared to designate the simulation’s termination condition. Unlike variables and constants, params have a known value at compile time and can affect how the program is compiled. In this code, we’ll use it to switch between tolerance-based and iteration-based termination.
With all that set up, we can move on to setting up our arrays.
Using 2D Arrays
In order to solve the 2D Poisson Equation in Chapel, we’ll need a way to represent the 2D quantities and . Like the NumPy arrays used in the tutorial, Chapel’s arrays can have an arbitrary number of dimensions, so we can continue using them in a similar manner to the previous post.
The syntax we used last time for declaring a 1D array:
var a: [0..<nx] real;
is generalized for 2D arrays as follows:
var a: [0..<ny, 0..<nx] real;
Here, a is defined over the outer product of all the indices in the two ranges.
Although declaring our and arrays using the above syntax would work, it will be useful to store the set of indices that defines our problem space using a named variable. This is partially a stylistic choice, but it also makes it easier to modify the code for multi-locale execution later on (covered in the next post in this series).
We start by declaring some named index sets (or [note:Chapel’s domains represent sets of indices. They can be used to define arrays (or in our case, groups of arrays that share the same index set) and can also be iterated over directly. See these docs for an introduction to domains.]):
|
|
Here, the list of ranges within the curly braces on the first line creates a domain that will represent the index set for our simulation. The expand method is used to create another domain, spaceInner, which represents the same indices, excluding those around the boundary
(i.e.,{1..<(nx-1), 1..<(ny-1)}). We’ll use this later for applying a stencil operation to p.
With our named domains defined, we can declare our 2D arrays as follows:
var p, b: [space] real;
The main procedure
Many programming languages use a main function to designate the program’s starting point. Like Python, Chapel doesn’t require this, but does support it. For this code, which is longer than the diffusion code, we’ll try to keep things organized by breaking our problem into multiple subroutines, putting the array declaration and setup into the main procedure. Without going into too much detail, this procedure makes use of the array declaration syntax we discussed above, then sets up initial and boundary conditions, and finally calls another procedure, solvePoisson that we’ll discuss below:
|
|
The Solver and where Clauses
The algorithm for solving the Poisson Equation is essentially the same as the Diffusion Equation; the only difference is that this code will support two different termination conditions. It will:
- create a temporary copy of
pcalledpn - do the following…
- swap
pandpn - apply the Poisson Equation to
pn, storing inp - apply the boundary conditions to
p
- swap
- …until the termination condition is met:
- tolerance: the relative change in the L1 norm is less than the given tolerance
- iteration: the specified number of iterations has run
For the tolerance condition, we have the following routine, which again breaks parts of the approach into helper procedures: poissonKernel and neumannBC:
|
|
Note that we’re still preventing the simulation from running for more than maxIters iterations to avoid looping forever in case the simulation cannot converge; however, with the default simulation constants, this code will terminate well before maxIters is reached.
more on + reduce
Here, we’re using Chapel’s native reductions to compute the relative change in p’s L1 norm from the previous iteration.
Focusing in on the denominator, abs(pn) applies the math function abs to pn in a promoted element-wise manner (see more on promotion here), creating an iterable stream of values. The reduce operator then reduces those values into a single number by summing them. This is a simple and performant way to get the sum of the absolute-value of pn. A similar process occurs in the numerator where the reduced expression is slightly more complex.
In addition to sum-reductions, Chapel supports a variety of others. Check out the docs for more information.
And for the iteration condition, we have a slightly simplified routine that uses the same helper procedures:
|
|
Notice that both procedures are named solvePoisson and accept the same arguments. Typically, defining two such procedures would be an error; however these routines use where clauses to allow the compiler to select which of the two is called, making the double-definition legal. When the program is compiled, the compiler looks at the value of useTolerance and uses the matching overload of solvePoisson.
Note that useTolerance is defined with the config keyword, so a command-line argument can be passed to the compiler with the -s or --set flag to change its value. For example, this command:
chpl nsPoisson.chpl --set useTolerance=false
would compile the program to use iteration-based termination.
The Poisson Kernel
The poissonKernel procedure used by both of the above routines applies the discretized Poisson Equation to p. The equation is rearranged to fit our algorithm:
Where the on the left corresponds to p, and each on the right corresponds to pn. In Chapel, we have the following:
|
|
Note that a forall loop is used, indicating to the compiler that this loop is order-independent and that spaceInner’s parallel iterator can be used to break up the work across multiple tasks. This permits the code to be executed in parallel on multi-core hardware. It’s also notable that nested loops are unnecessary even though we’re iterating over a 2D array’s indices because spaceInner’s iterator yields 2-tuples.
The solvePoisson procedures also call the following boundary condition function to apply Neumann boundary conditions to the top and bottom walls. The other boundary conditions don’t have to be applied each iteration because the elements around the boundary of p are never updated after being set:
|
|
Running and generating plots
To try the simulation out on your own machine, download and place the following three files in the same directory. The first is the full code we just went over, while the second has a helper procedure for writing p to a file and calling a Python plotting script (the third file).
nsPoisson.chpl
|
|
surfPlot.chpl
|
|
surfPlot.py
|
|
For the plotting to work, make sure you have python3 in your path with numpy and matplotlib available in your Python environment.
Run the following command to compile the program:
chpl nsPoisson.chpl --fast
And try running it with:
./nsPoisson --sourceMag=500 --makePlots=true
After the simulation finishes, the plotting script should generate the following figure (named solution.png, in the same directory as the plotting script). Try adjusting the config consts
(e.g., --sourceMag=-200) to see how they affect the results.
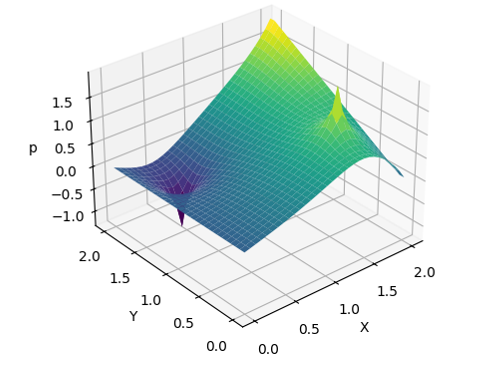
Poisson Solution
And with that, we have a full Poisson Equation solver that contains all the essential elements from Steps 9 and 10 of the Python tutorial. Not only that, it makes use of parallel iteration and occupies just under 100 lines of code. Because Chapel is a high-performance compiled language, we can also expect our code to get great performance. To put that to the test, let’s compare with a parallel implementation in another high performance compiled language that’s often used in scientific computing.
Performance Comparison with C++/OpenMP
[note:Caveat: I am not a C++/OpenMP expert, so there are certainly places where the performance, and perhaps readability, of the C++ code could be improved; however, the purpose of this exercise is to show how a straightforward Chapel implementation compares to a straightforward C++/OpenMP implementation without pulling out all the stops for either technology.] the Chapel code to C++, using OpenMP to parallelize when possible, we have the following source and header files:
nsPoisson.cpp
|
|
nsPoisson.h
|
|
Besides being longer, there are few notable differences with the Chapel code:
- argument parsing is done manually using a function in the utility file (shown below)
- in practice, one would likely use a third-party library for this
- 2D arrays are represented as nested vectors (i.e.,
vector<vector<double>>) - switching between tolerance- and iteration-based termination relies on an
#ifdefmacro- this is shorter than having two separate procedures but could be harder to read
- OpenMP pragmas are used to transform serial loops into parallel loops
Here are some other source files that will be useful for running this code yourself (note: make sure to use -DCMAKE_BUILD_TYPE=Release when invoking cmake):
utils.cpp
|
|
utils.h
|
|
CMakeLists.txt
|
|
Experiment Setup and Results:
Both the Chapel and C++ codes were compiled to run for 10,000 iterations (no tolerance termination) on various problem sizes. To keep the simulation numerically stable, the ratio of physical length per finite-difference point was kept constant at r=2.0/30. The programs were executed with the following arguments:
./nsPoisson --nx=<n> --ny=<n> --xLen=<n*r> --yLen=<n*r>
where n ranged from 64 to 2048 by powers of 2. So, the total number of points ranged from 4096 to just over four million.
Testing was performed on a Linux system with the following CPU specs: 2.8GHz 32-Core AMD EPYC, 256 MiB L3 cache, hyper-threading enabled. GCC 7.5 was used to compile the C++ code, and it was executed with OMP_PROC_BIND=true and OMP_NUM_THREADS=32. The Chapel codes were compiled with version 2.0 using the default linux64 configuration. Execution times were computed as the average of three trials.
This experiment produced the following results:
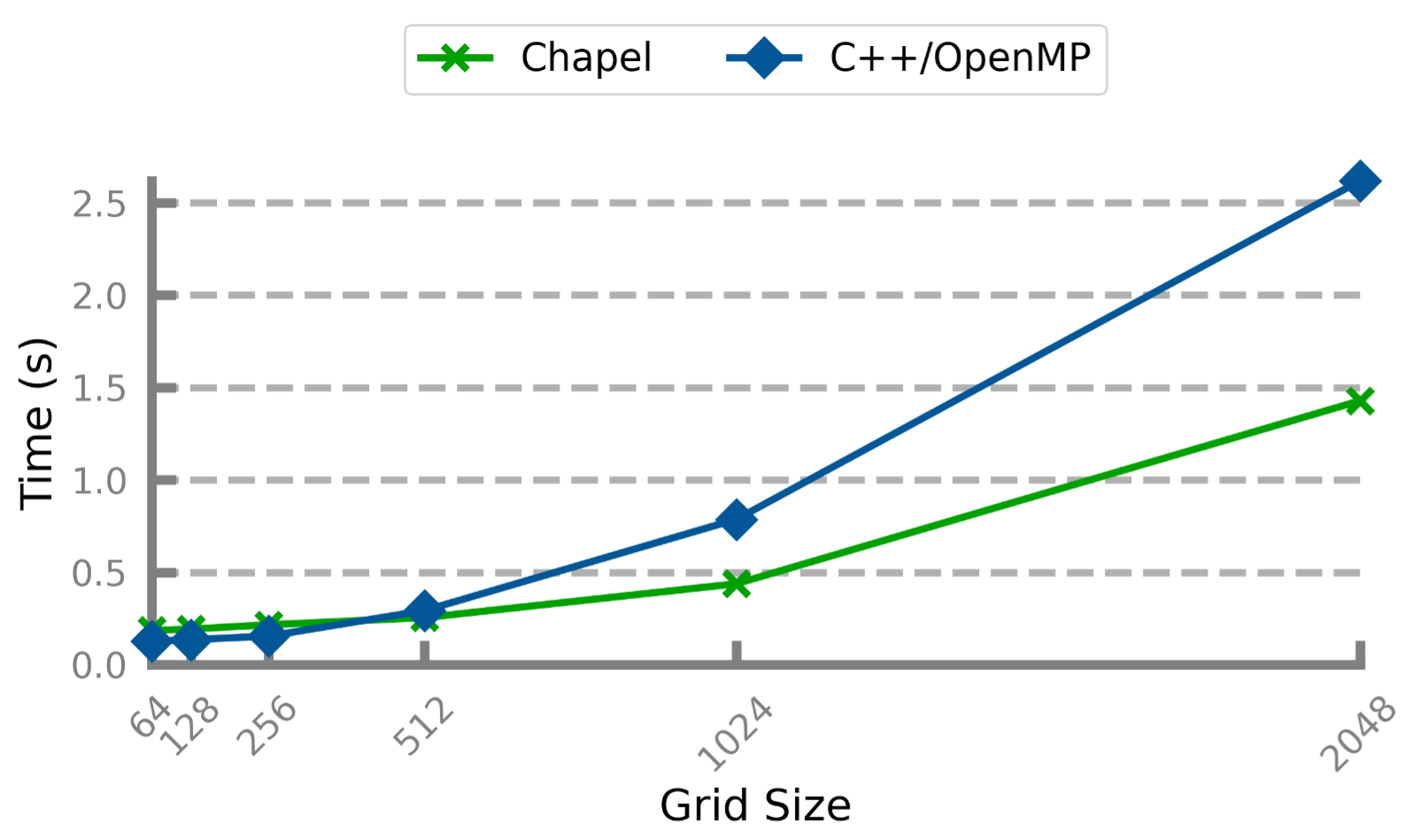
Performance Comparison: Chapel & C++
For smaller problem sizes, the Chapel and C++ code perform similarly, with C++ coming out slightly ahead. As the problem size increases, Chapel begins to [note:This could be because the C++ code uses nested vectors to represent the 2D arrays, meaning that the outer vector contains pointers to the inner vectors (which, in general, are not allocated near each other on the heap). As such, cache misses are potentially more likely as compared to the Chapel code whose 2D arrays allocate all the data in one contiguous block in memory. For larger problems, where the entire array cannot fit in cache, this effect may be harming the C++ code’s performance more significantly. To alleviate this, the logical 2D arrays could instead be implemented as 1D vectors with length nx * ny.] by a notable margin.
To put the above results in perspective, we also looked at the performance of the Python code run on the same hardware with the same problem sizes. This included a normal Python script (essentially copied from the tutorial), and a [note:Again, I’m not an expert with Numba or other Python parallelization techniques. It’s certainly possible that one could squeeze more performance out of the Python code with more effort and expertise. It is also possible to run the code on an accelerated build of NumPy that makes use of OpenMP-enabled BLAS or various vendor-accelerated backends; however, despite a significant amount of effort, I was unable to procure such a build on my testing hardware.] of the same script which uses the @stencil and @jit(nopython=True, parallel=True) annotations to improve performance.
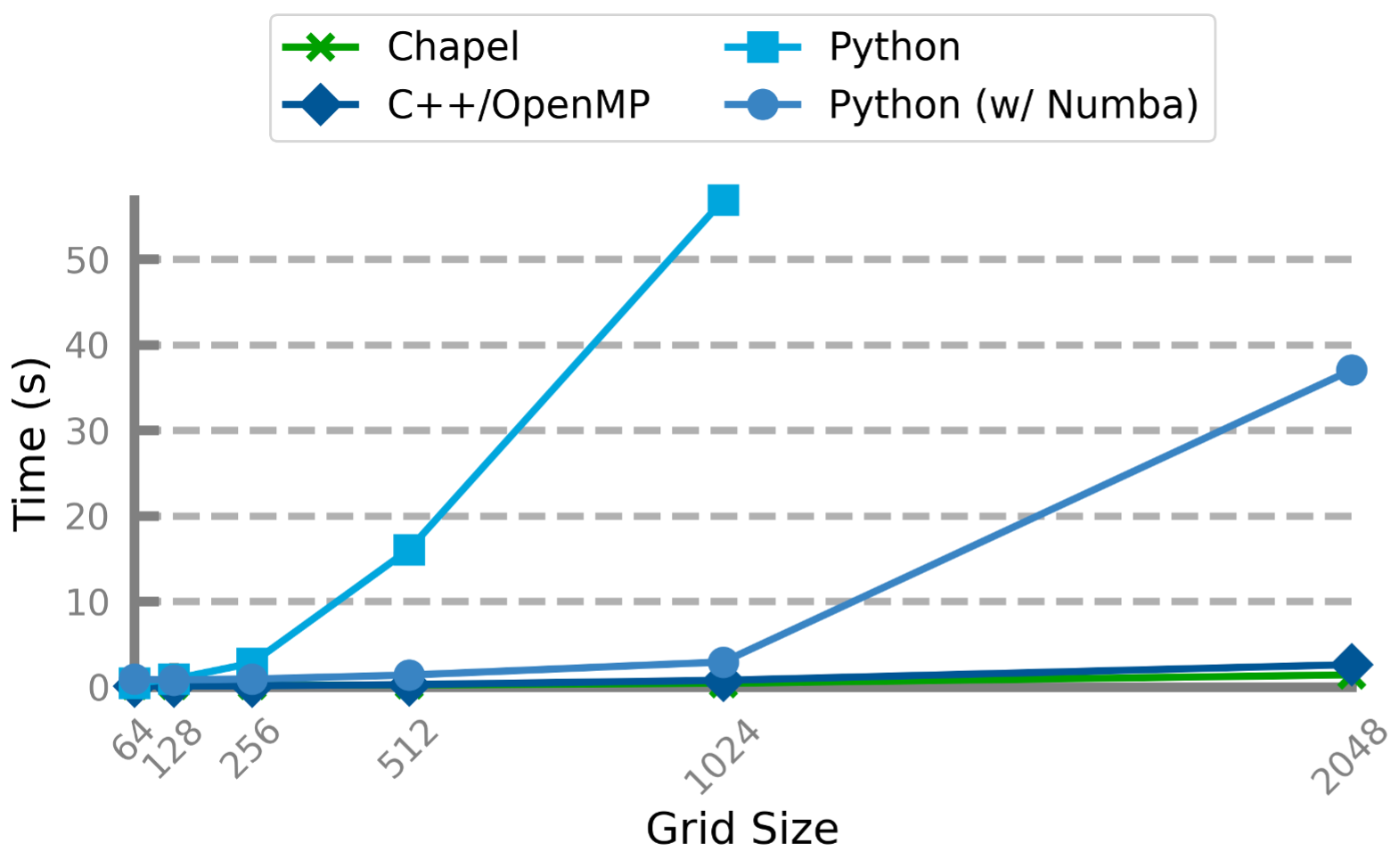
Performance Comparison: Chapel, C++, Python
Here, as you may expect, pure Python is slower than the compiled languages by orders of magnitude on the larger problem sizes. Numba is able to produce competitive performance up to a point, but slows down significantly for the largest problem size.
Conclusion
This article aimed to use a common scientific computation to show that Chapel’s productive syntax (including its powerful multidimensional arrays) and its excellent performance (powered by its first class parallel computing features) make it a great candidate for use in scientific computing. Not only does it fit steps 9 and 10 of the 12 Steps to Navier-Stokes tutorial — with helpful command-line features for configuring the simulation — into about 100 lines of code; it’s also able to outperform compiled Python and Parallel C++.
In this series’ next post, we’ll dive into Chapel’s distributed programming features and see how easy it is to modify the Poisson solver to run across multiple compute nodes. After that, we’ll continue stepping up the complexity of our computations by looking at Step 11 of the tutorial — a full cavity-flow Navier-Stokes simulation. This simulation will also be able to run on a multi-node cluster or supercomputer, and to show that Chapel’s productivity doesn’t come with a performance penalty, we’ll compare its performance with a C++/MPI/OpenMP version of the same code.
The Chapel Team would like to express its sincere gratitude towards the Barba Group for creating and maintaining the CFD Python tutorials.
